PDK#
gdsfactory includes a generic Process Design Kit (PDK), that you can leverage to create your own.
A process design kit (PDK) includes:
LayerStack: different layers with different thickness, z-position, materials and colors.
Design rule checking deck DRC: Manufacturing rules capturing min feature size, min spacing … for the process.
A library of Parametric cells or fixed cells (no parameters).
The PDK allows you to register:
cellparametric cells that return Components from a ComponentSpec (string, Component, ComponentFactory or dict). Also known as parametric cell functions.cross_sectionfunctions that return CrossSection from a CrossSection Spec (string, CrossSection, CrossSectionFactory or dict).layersthat return a GDS Layer (gdslayer, gdspurpose) from a string, an int or a Tuple[int, int].
Thanks to activating a PDK you can access components, cross_sections or layers using a string, a function or a dict.
Depending on the active pdk:
get_layerreturns a Layer from the registered layers.get_componentreturns a Component from the registered cells.get_cross_sectionreturns a CrossSection from the registered cross_sections.
Layers#
GDS layers are defined as a tuple of two integer numbers gdslayer/gdspurpose
You can define all the layers using your PDK:
import pathlib
from functools import partial
import pytest
from pytest_regressions.data_regression import DataRegressionFixture
import gdsfactory as gf
from gdsfactory.difftest import difftest
from gdsfactory.technology import (
LayerMap,
)
from gdsfactory.typings import Layer
gf.gpdk.PDK.activate()
nm = 1e-3
class LayerMapDemo(LayerMap):
WG: Layer = (1, 0)
DEVREC: Layer = (68, 0)
PORT: Layer = (1, 10)
PORTE: Layer = (1, 11)
LABEL_INSTANCES: Layer = (206, 0)
LABEL_SETTINGS: Layer = (202, 0)
LUMERICAL: Layer = (733, 0)
M1: Layer = (41, 0)
M2: Layer = (45, 0)
M3: Layer = (49, 0)
N: Layer = (20, 0)
NP: Layer = (22, 0)
NPP: Layer = (24, 0)
OXIDE_ETCH: Layer = (6, 0)
P: Layer = (21, 0)
PDPP: Layer = (27, 0)
PP: Layer = (23, 0)
PPP: Layer = (25, 0)
PinRec: Layer = (1, 10)
PinRecM: Layer = (1, 11)
SHALLOWETCH: Layer = (2, 6)
SILICIDE: Layer = (39, 0)
SIM_REGION: Layer = (100, 0)
SITILES: Layer = (190, 0)
SLAB150: Layer = (2, 0)
SLAB150CLAD: Layer = (2, 9)
SLAB90: Layer = (3, 0)
SLAB90CLAD: Layer = (3, 1)
SOURCE: Layer = (110, 0)
TE: Layer = (203, 0)
TEXT: Layer = (66, 0)
TM: Layer = (204, 0)
Text: Layer = (66, 0)
VIA1: Layer = (44, 0)
VIA2: Layer = (43, 0)
VIAC: Layer = (40, 0)
WGCLAD: Layer = (111, 0)
WGN: Layer = (34, 0)
WGclad_material: Layer = (36, 0)
LAYER = LayerMapDemo
some generic components use some
Layer |
Purpose |
|---|---|
LABEL_INSTANCE |
for adding instance labels on |
MTOP |
for top metal routing |
class LayersConvenient(LayerMap):
LABEL_INSTANCE: Layer = (66, 0)
Cross_Sections#
You can create a CrossSection from scratch or you can customize the cross_section functions in gf.cross_section
from gdsfactory.cross_section import CrossSection, cross_section, xsection
from gdsfactory.typings import LayerSpec
@xsection
def strip2(
width: float = 0.5,
layer: LayerSpec = (2, 0),
radius: float = 10.0,
radius_min: float = 5,
**kwargs,
) -> CrossSection:
"""Return Strip cross_section."""
return cross_section(
width=width,
layer=layer,
radius=radius,
radius_min=radius_min,
**kwargs,
)
c = gf.components.straight(cross_section=strip2)
c.plot()

@xsection
def pin(
width: float = 0.5,
layer: LayerSpec = "WG",
radius: float = 10.0,
radius_min: float = 5,
layer_p: LayerSpec = (21, 0),
layer_n: LayerSpec = (20, 0),
width_p: float = 2,
width_n: float = 2,
offset_p: float = 1,
offset_n: float = -1,
**kwargs,
) -> CrossSection:
"""Return PIN cross_section."""
sections = (
gf.Section(layer=layer_p, width=width_p, offset=offset_p),
gf.Section(layer=layer_n, width=width_n, offset=offset_n),
)
return cross_section(
width=width,
layer=layer,
radius=radius,
radius_min=radius_min,
sections=sections,
**kwargs,
)
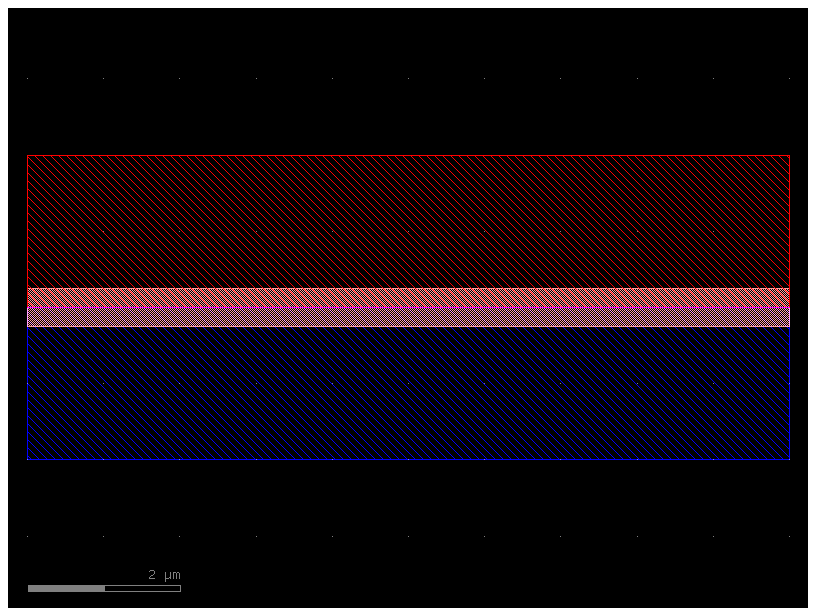
c = gf.components.straight(cross_section=pin)
c.plot()

@xsection
def strip_wide(
width: float = 3,
layer: LayerSpec = (2, 0),
radius: float = 10.0,
radius_min: float = 5,
**kwargs,
) -> CrossSection:
"""Return Strip cross_section."""
return cross_section(
width=width,
layer=layer,
radius=radius,
radius_min=radius_min,
**kwargs,
)
strip = gf.cross_section.strip
cross_sections = dict(strip_wide=strip_wide, pin=pin, strip=strip)
Cells#
Cells are functions that return components. They are parametrized and accept other cells as parameters, so that you can build many levels of complexity. Cells are also known as PCells or parametric cells.
You can customize the function’s default arguments easily thanks to functools.partial
Let us now customize the default arguments of a library of cells.
For example, you can make some wide MMIs for a particular technology. Let us now assume that the best MMI width you found to be 9um.
def mmi1x2(width_mmi: float = 9, **kwargs) -> gf.Component:
c = gf.components.mmi1x2(width_mmi=width_mmi)
return c
def mmi2x2(width_mmi: float = 9, **kwargs) -> gf.Component:
c = gf.components.mmi2x2(width_mmi=width_mmi)
return c
cells = dict(mmi1x2=mmi1x2, mmi2x2=mmi2x2)
You can define a new PDK by creating a function that customizes partial parameters of the generic functions.
Assuming this PDK uses layer (41, 0) for the pads (instead of the layer used in the generic pad function):
pad_custom_layer = partial(gf.components.pad, layer=(41, 0))
c = pad_custom_layer()
c.plot()

PDK#
You can register Layers, ComponentFactories (Parametric cells) and CrossSectionFactories (cross_sections) into a PDK.
Then you can access them through a string after you activate the pdk PDK.activate().
LayerSpec#
You can access layers of the active PDK using the layer name, a tuple/list of two numbers or a layer from a LayerMap object (such as LAYER.WG).
from gdsfactory.gpdk import get_generic_pdk
generic_pdk = get_generic_pdk()
pdk1 = gf.Pdk(
name="fab1",
layers=LAYER,
cross_sections=cross_sections,
cells=cells,
layer_views=generic_pdk.layer_views,
)
pdk1.activate()
pdk1.get_layer("WG")
<LayerMapDemo.WG: 1>
pdk1.get_layer((1, 0))
1
CrossSectionSpec#
You can access cross_sections of the pdk from the cross_section name, or using a dict to customize the cross-section.
pdk1.get_cross_section("pin")
CrossSection(sections=(Section(width=0.5, offset=0.0, insets=None, layer='WG', port_names=('o1', 'o2'), port_types=('optical', 'optical'), name='_default', hidden=False, simplify=None, skip_transition=False, width_function=None, offset_function=None), Section(width=2.0, offset=1.0, insets=None, layer=(21, 0), port_names=(None, None), port_types=('optical', 'optical'), name='s_24eda351', hidden=False, simplify=None, skip_transition=False, width_function=None, offset_function=None), Section(width=2.0, offset=-1.0, insets=None, layer=(20, 0), port_names=(None, None), port_types=('optical', 'optical'), name='s_4e8950a1', hidden=False, simplify=None, skip_transition=False, width_function=None, offset_function=None)), components_along_path=(), radius=10.0, radius_min=5.0, bbox_layers=None, bbox_offsets=None)
cross_section_spec_string = "pin"
c = gf.components.straight(cross_section=cross_section_spec_string)
c.plot()

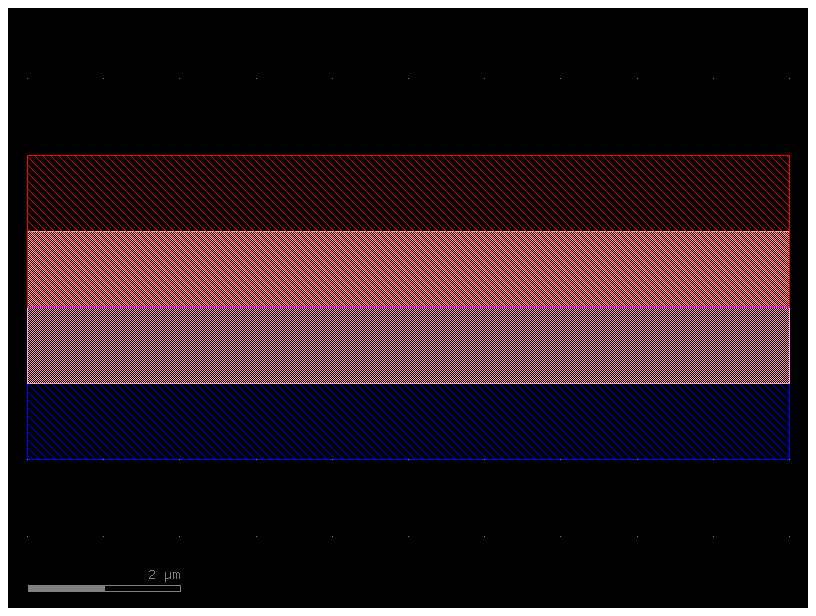
xs = gf.get_cross_section("pin", width=2)
wg_pin = gf.components.straight(cross_section=xs)
wg_pin.plot()
ComponentSpec#
You can get a component of the active pdk using the cell name (string) or a dict.
c = pdk1.get_component("mmi1x2")
c.plot()
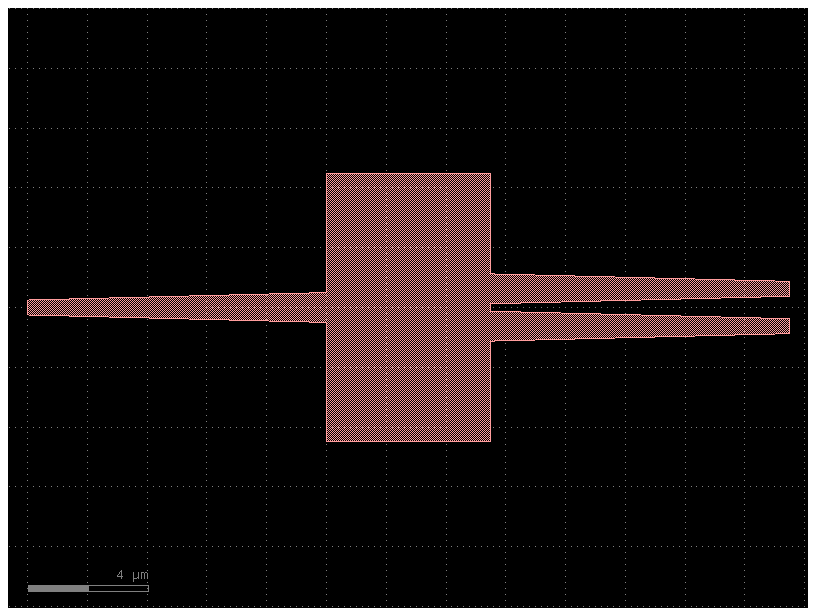
c = pdk1.get_component(dict(component="mmi1x2", settings=dict(length_mmi=10)))
c.plot()

Layer Views#
Layer Views represent the stipples and colors used to render the layers in Klayout.
You can define the layer views
From a Klayout
lyp(layer properties file).From a
yamlfile.From scratch, adding all your layers into a class.
Now, let us generate the layers definition code from a KLayout lyp file.
from gdsfactory.config import PATH
import gdsfactory as gf
LAYER_VIEWS = gf.technology.LayerViews(PATH.klayout_lyp)
LYP files are defined in XML and are only easy to edit in the KLayout Graphical User Interface (GUI).
Sometimes you also want to maintain your layer stipple and colors in YAML.
LAYER_VIEWS = gf.technology.LayerViews(PATH.klayout_yaml)
PDK#
To link layout and circuit models to a PDK you need to register them in a PDK.
There are also many other pdk settings you can define in the PDK.
PDK.namePDK.versionPDK.cellsfunctions registered in the PDK to create parametric cells.PDK.cross_sectionsfunctions registered in the PDK to create cross_sections.PDK.layerslayers registered in the PDK.PDK.layer_stack3D layer stack registered in the PDK. Useful for device level simulations.PDK.layer_viewsto define how to render layer colors and styles in KLayout viewer.PDK.layer_transitionsto define how to transition between layers automatically.PDK.materials_indexto define the materials index for device level simulations.PDK.constantsto define PDK constants.PDK.connectivityto define connectivity between layers. For metal connectivity traceability.
import gdsfactory as gf
from functools import partial
from gdsfactory.config import PATH, __version__
from gdsfactory.cross_section import cross_sections
from gdsfactory.gpdk.simulation_settings import materials_index
from gdsfactory.get_factories import get_cells
from gdsfactory.pdk import Pdk
from gdsfactory.gpdk.layer_stack import LAYER_STACK, LAYER
from gdsfactory.technology import LayerViews
LAYER_VIEWS = LayerViews(filepath=PATH.klayout_yaml)
LAYER_CONNECTIVITY = [
("NPP", "VIAC", "M1"),
("PPP", "VIAC", "M1"),
("M1", "VIA1", "M2"),
("M2", "VIA2", "M3"),
]
cells = get_cells([gf.components])
containers_dict = get_cells([gf.containers])
layer_transitions = {
LAYER.WG: partial(gf.c.taper, cross_section="strip", length=10),
(LAYER.WG, LAYER.WGN): "taper_sc_nc",
(LAYER.WGN, LAYER.WG): "taper_nc_sc",
LAYER.M3: "taper_electrical",
}
class GenericConstants(gf.Constants):
"""Generic PDK constants."""
fiber_input_to_output_spacing: float = 200.0
metal_spacing: float = 10.0
pad_pitch: float = 100.0
pad_size: tuple[float, float] = (80.0, 80.0)
wavelength: float = 1.55
PDK = Pdk(
name="generic",
version=__version__,
cells={c: cells[c] for c in cells if c not in containers_dict},
containers=containers_dict,
cross_sections=cross_sections,
layers=LAYER,
layer_stack=LAYER_STACK,
layer_views=LAYER_VIEWS,
layer_transitions=layer_transitions, # How to transition between layers.
materials_index=materials_index, # Material index for device level simulations.
constants=GenericConstants(),
connectivity=LAYER_CONNECTIVITY, # For tracing connectivity across layers such as metals.
)
Testing PDK cells#
To make sure all your PDK PCells produce the components that you want, it is important to test your PDK cells.
As you write your own cell functions you want to make sure you do not break or produce unwanted regressions later on. You should write tests for this.
Make sure you create a test_components.py file for pytest to test your PDK. See for example the tests in the ubc PDK
Pytest-regressions automatically creates the CSV and YAML files for you, gdsfactory.gdsdiff will store the reference GDS in ref_layouts and check for geometry differences using XOR.
gdsfactory is NOT backwards compatible, which means that the package will keep improving and evolving.
To make your work stable you should install a specific version and pin the version in your
requirements.txtorpyproject.tomlasgdsfactory==9.43.0replacing9.43.0by whatever version you end up using.Before you upgrade gdsfactory to a newer version make sure your tests pass to make sure that things behave as expected
"""This code tests all your cells in the PDK
It will test:
1. difftest: Will test the GDS geometry of a new GDS compared to a reference.
2. settings test: Will compare the settings in YAML with a reference YAML.
"""
try:
dirpath = pathlib.Path(__file__).absolute().with_suffix(".gds")
except Exception:
dirpath = pathlib.Path.cwd()
component_names = list(pdk1.cells.keys())
factory = pdk1.cells
@pytest.fixture(params=component_names, scope="function")
def component_name(request) -> str:
return request.param
def test_gds(component_name: str) -> None:
"""Avoid regressions in GDS files. Runs XOR and computes the area."""
component = factory[component_name]()
test_name = f"fabc_{component_name}"
difftest(component, test_name=test_name, dirpath=dirpath)
def test_settings(component_name: str, data_regression: DataRegressionFixture) -> None:
"""Avoid regressions in component settings and ports."""
component = factory[component_name]()
data_regression.check(component.to_dict())
Compare gds files#
You can use the command line gf gds diff gds1.gds gds2.gds to overlay gds1.gds and gds2.gds files and show them in KLayout.
For example, if you changed the mmi1x2 and made it 5um longer by mistake, you could gf gds diff ref_layouts/mmi1x2.gds run_layouts/mmi1x2.gds and see the GDS differences in Klayout.
help(gf.diff)
Help on function diff in module gdsfactory.difftest:
diff(ref_file: pathlib.Path | str, run_file: pathlib.Path | str, xor: bool = True, test_name: str = '', ignore_sliver_differences: bool | None = None, ignore_cell_name_differences: bool | None = None, ignore_label_differences: bool | None = None, show: bool = False, stagger: bool = True, out_file: pathlib.Path | str | None = None, sliver_tolerance: int = 1) -> bool
Returns True if files are different, prints differences and shows them in klayout.
Args:
ref_file: reference (old) file.
run_file: run (new) file.
xor: runs xor on every layer between old and run files.
test_name: prefix for the new cell.
ignore_sliver_differences: if True, ignores any sliver differences in the XOR result. If None (default), defers to the value set in CONF.difftest_ignore_sliver_differences
ignore_cell_name_differences: if True, ignores any cell name differences. If None (default), defers to the value set in CONF.difftest_ignore_cell_name_differences
ignore_label_differences: if True, ignores any label differences when run in XOR mode. If None (default) defers to the value set in CONF.difftest_ignore_label_differences
show: shows diff in klayout.
stagger: if True, staggers the old/new/xor views. If False, all three are overlaid.
out_file: if not None, saves the diff to the specified file.
sliver_tolerance: tolerance in database units for sliver detection. Default is 1.
mmi1 = gf.components.mmi1x2(length_mmi=5)
mmi2 = gf.components.mmi1x2(length_mmi=6)
gds1 = mmi1.write_gds()
gds2 = mmi2.write_gds()
gf.diff(gds1, gds2)
cell name differs mmi1x2_gdsfactorypcomponentspmmispmmi1x2_WNone_WT1_LT10_85778fc7
Running XOR on differences...
: XOR difference on layer WG (1/0)
True

